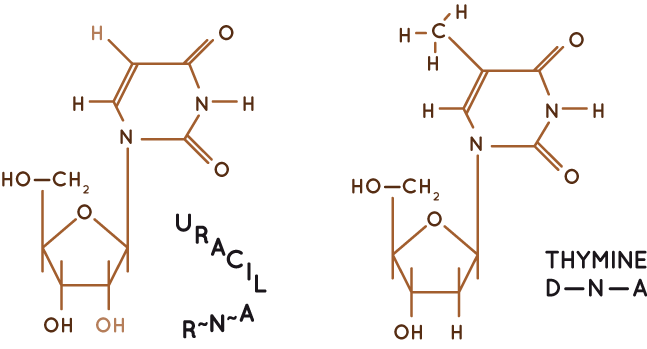
↗ RNA distinguishes itself from DNA by its sensitivity towards alkaline caused by an additional OH-group on the ribose.
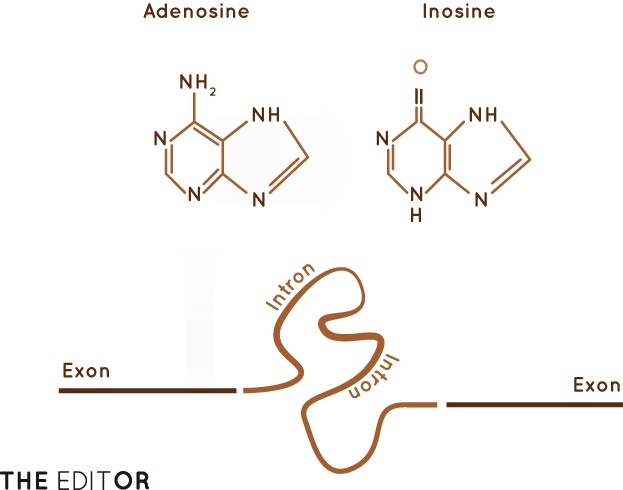
↗ Detailed chemical analyses revealed that RNA shares three bases with DNA: adenine, cytosine and guanine. In contrast, uracil is unique to RNA whereas thymine is generally present in DNA.
↗ Nucleic acids were isolated from various organisms.
↗ The RNA building blocks ATP and GTP were proposed to be the cell’s general energy source: life’s engine.